 
(2) Artificial Selection-motivated by these biological results, we examine the algorithmic problem of designing the most stable and unstable mRNA sequences which code for a target protein. 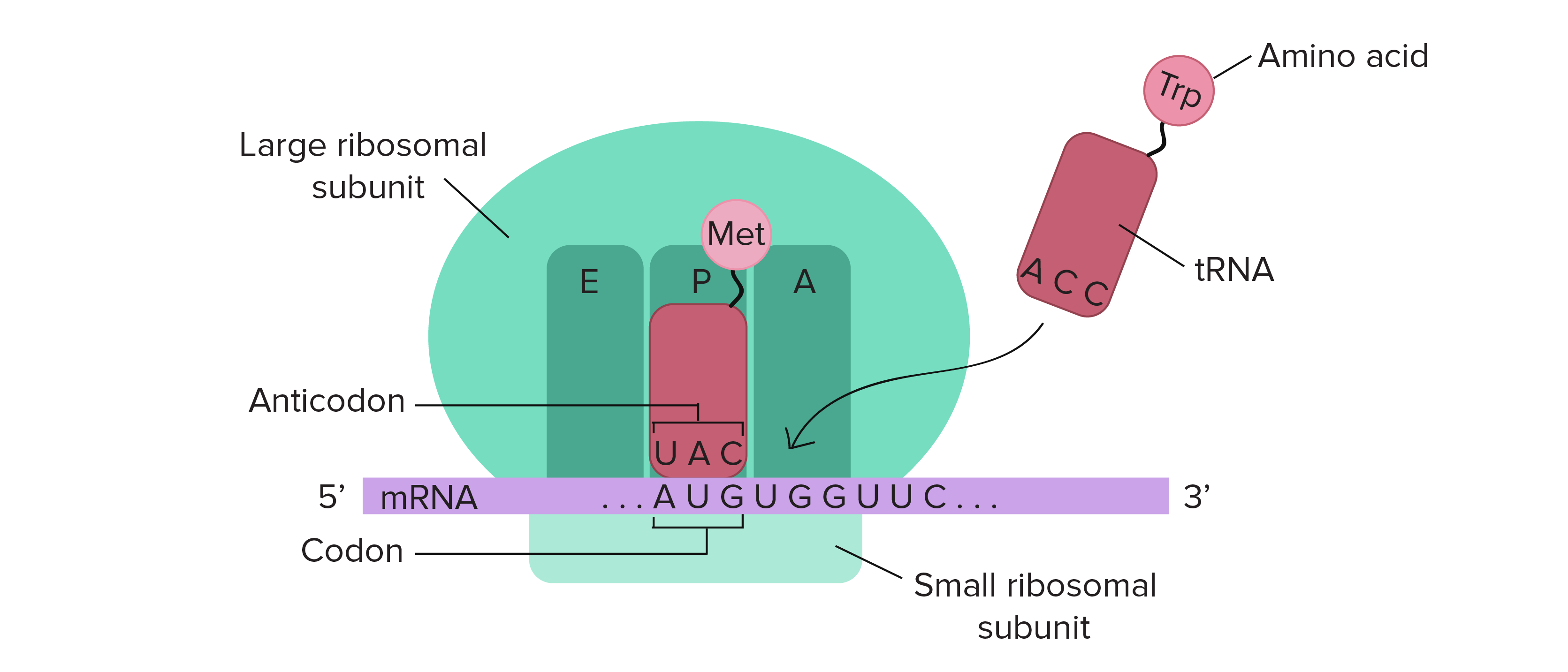
This suggests that the stability of mRNA sequences is subject to natural selection. We provide evidence that in all genomic structures highly stable sequences are disproportionately abundant, and in 19 of 36 cases highly unstable sequences are disproportionately abundant. This experiment was conducted on over 27,000 sequences from 34 microbial species with 36 genomic structures. In particular: (1) Natural Selection-we perform the first large-scale computational experiment comparing the stability of mRNA sequences from a variety of organisms to random synonymous sequences which respect the codon preferences of the organism. In this paper, we explore how nature takes advantage of this freedom and how to algorithmically design structures more energetically favorable than have been built through natural selection. Thus nature has wide latitude to select among mRNA sequences which are informationally equivalent, but structurally and energetically divergent. This freedom implies that the number of possible RNA sequences coding for a given protein grows exponentially in the length of the protein. Certain residues (e.g., tryptophan) have only a single corresponding codon, while other residues (e.g., arginine) have as many as six corresponding codons. Because there are more codons than residues, there is inherent redundancy in the coding. Messenger RNA (mRNA) sequences serve as templates for proteins according to the triplet code, in which each of the 4(3) = 64 different codons (sequences of three consecutive nucleotide bases) in RNA either terminate transcription or map to one of the 20 different amino acids (or residues) which build up proteins. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
